Structural basis for immune cell binding of Fusobacterium nucleatum via the trimeric autotransporter adhesin CbpF
Marongiu, G.L., Fink, U., Oder, A., von Kries, J.P., Roderer, D.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cell surface protein | 489 | Fusobacterium nucleatum | Mutation(s): 0 Gene Names: FN1499 | 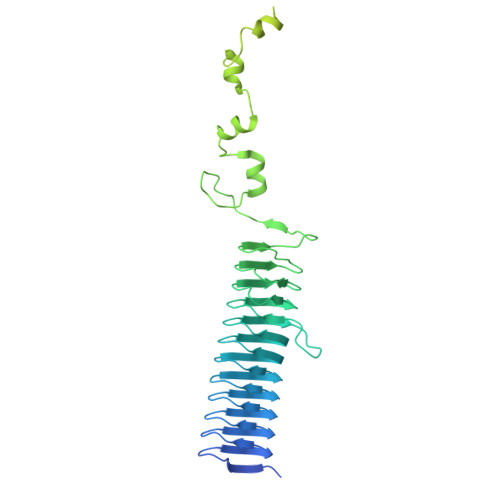 | |
UniProt | |||||
Find proteins for Q8RIS0 (Fusobacterium nucleatum subsp. nucleatum (strain ATCC 25586 / DSM 15643 / BCRC 10681 / CIP 101130 / JCM 8532 / KCTC 2640 / LMG 13131 / VPI 4355)) Explore Q8RIS0 Go to UniProtKB: Q8RIS0 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q8RIS0 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Carcinoembryonic antigen-related cell adhesion molecule 1 | 400 | Homo sapiens | Mutation(s): 0 Gene Names: CEACAM1, BGP, BGP1 | 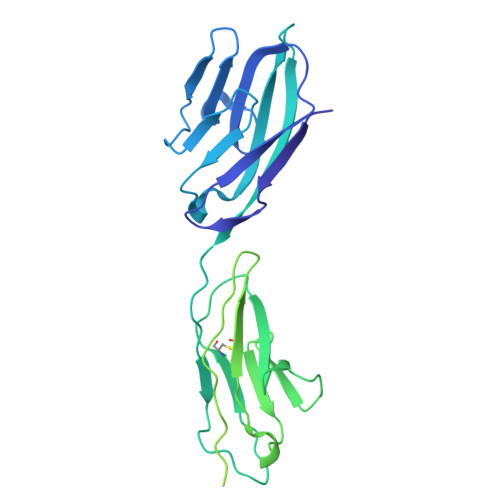 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P13688 (Homo sapiens) Explore P13688 Go to UniProtKB: P13688 | |||||
PHAROS: P13688 GTEx: ENSG00000079385 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P13688 | ||||
Glycosylation | |||||
| Glycosylation Sites: 6 | Go to GlyGen: P13688-1 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAG Query on NAG | G [auth D] H [auth D] I [auth D] J [auth D] K [auth D] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| RECONSTRUCTION | cryoSPARC | 4.4.0 |
| MODEL REFINEMENT | PHENIX | 1.21-5207 |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Leibniz Association | Germany | -- |