Cryo-EM structure of GPCR16-miniGs complex
Wang, X., Wu, L.J., Hua, T., Liu, Z.J.To be published.
Experimental Data Snapshot
Starting Model: in silico
View more details
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Guanine nucleotide-binding protein G(s) subunit alpha isoforms short,Guanine nucleotide-binding protein G(s) subunit alpha isoforms short,Guanine nucleotide-binding protein G(t) subunit alpha-3 | 264 | Homo sapiens | Mutation(s): 0 Gene Names: GNAS, GNAS1, GSP EC: 3.6.5 |  | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P63092 (Homo sapiens) Explore P63092 Go to UniProtKB: P63092 | |||||
PHAROS: P63092 GTEx: ENSG00000087460 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P63092 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 | 366 | Homo sapiens | Mutation(s): 0 Gene Names: GNB1 |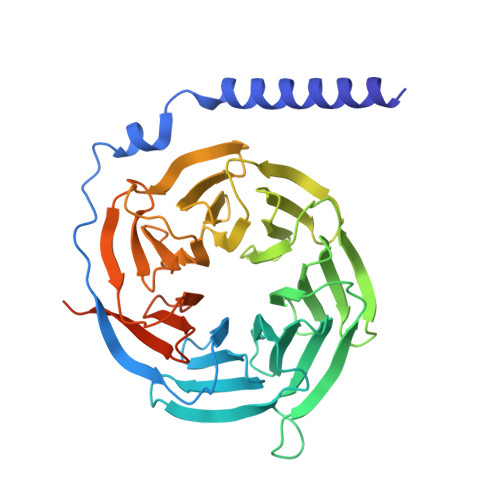  | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P62873 (Homo sapiens) Explore P62873 Go to UniProtKB: P62873 | |||||
PHAROS: P62873 GTEx: ENSG00000078369 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P62873 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 | C [auth G] | 71 | Homo sapiens | Mutation(s): 0 Gene Names: GNG2 |  |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P59768 (Homo sapiens) Explore P59768 Go to UniProtKB: P59768 | |||||
PHAROS: P59768 GTEx: ENSG00000186469 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P59768 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Nanobody 35 | D [auth N] | 135 | Homo sapiens | Mutation(s): 0 |  |
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Fusion protein 1,exo-alpha-sialidase,Taste receptor type 2 member 16,Fusion protein 2,exo-alpha-sialidase,Taste receptor type 2 member 16 | E [auth R] | 1,011 | Homo sapiens, Streptococcus pneumoniae | Mutation(s): 0 EC: 3.2.1.18 |  |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for A0A4J1WX37 (Streptococcus pneumoniae) Explore A0A4J1WX37 Go to UniProtKB: A0A4J1WX37 | |||||
Find proteins for Q9NYV7 (Homo sapiens) Explore Q9NYV7 Go to UniProtKB: Q9NYV7 | |||||
PHAROS: Q9NYV7 GTEx: ENSG00000128519 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Groups | A0A4J1WX37Q9NYV7 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| SA0 (Subject of Investigation/LOI) Query on SA0 | F [auth R] | 2-(hydroxymethyl)phenyl beta-D-glucopyranoside C13 H18 O7 NGFMICBWJRZIBI-UJPOAAIJSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Chinese Academy of Sciences | China | XDB37030104 |