Automated Structure- and Sequence-Based Design of Proteins for High Bacterial Expression and Stability.
Goldenzweig, A., Goldsmith, M., Hill, S.E., Gertman, O., Laurino, P., Ashani, Y., Dym, O., Unger, T., Albeck, S., Prilusky, J., Lieberman, R.L., Aharoni, A., Silman, I., Sussman, J.L., Tawfik, D.S., Fleishman, S.J.(2016) Mol Cell 63: 337-346
- PubMed: 27425410
- DOI: https://doi.org/10.1016/j.molcel.2016.06.012
- Primary Citation of Related Structures:
5HQ3 - PubMed Abstract:
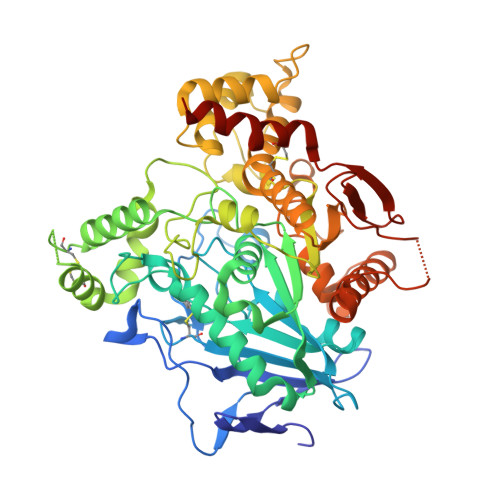
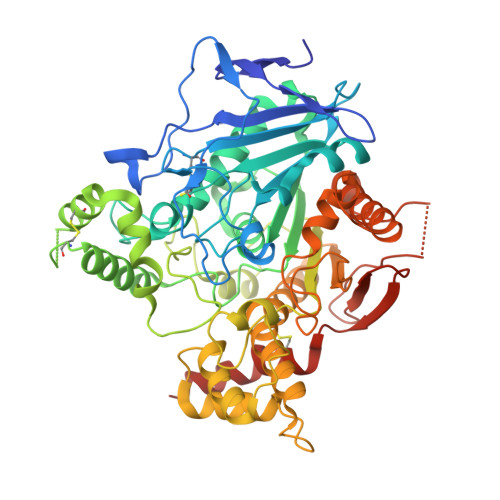
Upon heterologous overexpression, many proteins misfold or aggregate, thus resulting in low functional yields. Human acetylcholinesterase (hAChE), an enzyme mediating synaptic transmission, is a typical case of a human protein that necessitates mammalian systems to obtain functional expression. We developed a computational strategy and designed an AChE variant bearing 51 mutations that improved core packing, surface polarity, and backbone rigidity. This variant expressed at ~2,000-fold higher levels in E.?coli compared to wild-type hAChE and exhibited 20°C higher thermostability with no change in enzymatic properties or in the active-site configuration as determined by crystallography. To demonstrate broad utility, we similarly designed four other human and bacterial proteins. Testing at most three designs per protein, we obtained enhanced stability and/or higher yields of soluble and active protein in E.?coli. Our algorithm requires only a 3D structure and several dozen sequences of naturally occurring homologs, and is available at http://pross.weizmann.ac.il.
Organizational Affiliation:
Department of Biomolecular Sciences, Weizmann Institute of Science, Rehovot 7610001, Israel.