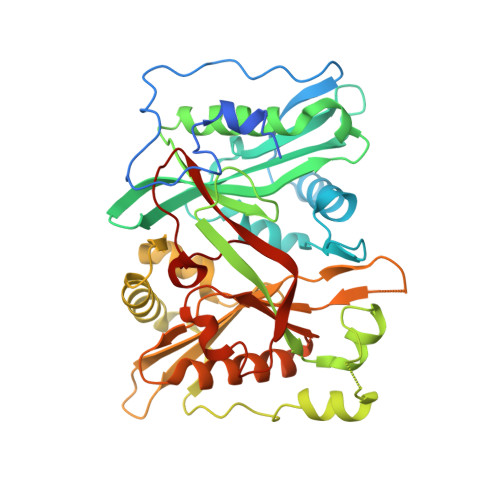
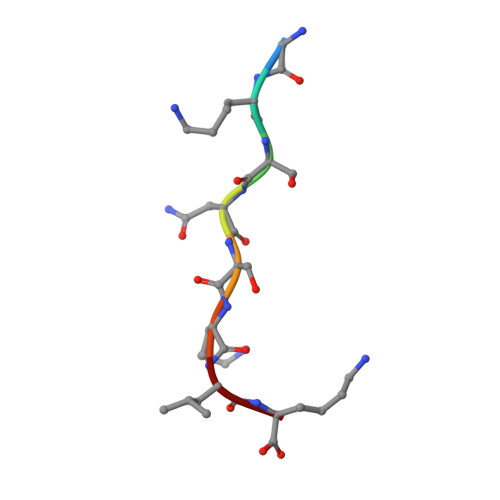
Structural and genomic decoding of human and plant myristoylomes reveals a definitive recognition pattern.
Castrec, B., Dian, C., Ciccone, S., Ebert, C.L., Bienvenut, W.V., Le Caer, J.P., Steyaert, J.M., Giglione, C., Meinnel, T.(2018) Nat Chem Biol 14: 671-679
- PubMed: 29892081
- DOI: https://doi.org/10.1038/s41589-018-0077-5
- Primary Citation of Related Structures:
5O9S, 5O9T, 5O9U, 5O9V - PubMed Abstract:
An organism's entire protein modification repertoire has yet to be comprehensively mapped. N-myristoylation (MYR) is a crucial eukaryotic N-terminal protein modification. Here we mapped complete Homo sapiens and Arabidopsis thaliana myristoylomes. The crystal structures of human modifier NMT1 complexed with reactive and nonreactive target-mimicking peptide ligands revealed unexpected binding clefts and a modifier recognition pattern. This information allowed integrated mapping of myristoylomes using peptide macroarrays, dedicated prediction algorithms, and in vivo mass spectrometry. Global MYR profiling at the genomic scale identified over a thousand novel, heterogeneous targets in both organisms. Surprisingly, MYR involved a non-negligible set of overlapping targets with N-acetylation, and the sequence signature marks for a third proximal acylation-S-palmitoylation-were genomically imprinted, allowing recognition of sequences exhibiting both acylations. Together, the data extend the N-end rule concept for Gly-starting proteins to subcellular compartmentalization and reveal the main neighbors influencing protein modification profiles and consequent cell fate.
Organizational Affiliation:
Institute for Integrative Biology of the Cell (I2BC), CEA, CNRS, Univ. Paris-Sud, Université Paris Saclay, Gif-sur-Yvette cedex, France.