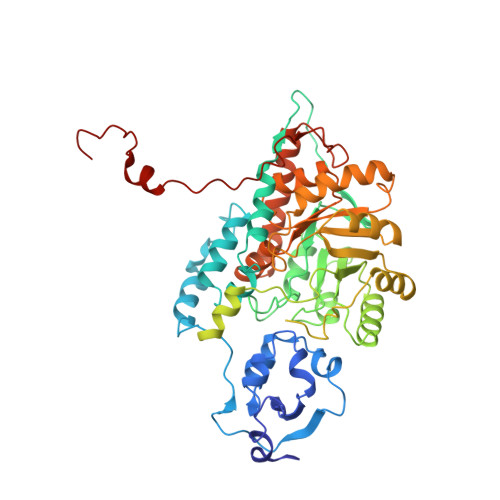
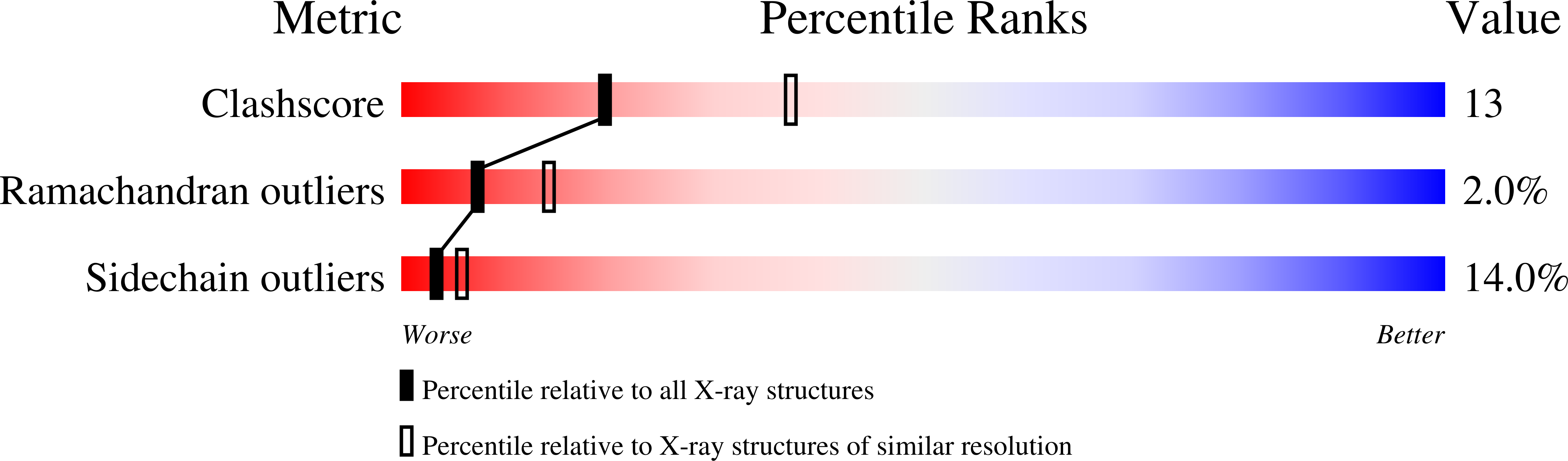The 2.6-A refined structure of the Escherichia coli recombinant Saccharomyces cerevisiae flavocytochrome b2-sulfite complex.
Tegoni, M., Cambillau, C.(1994) Protein Sci 3: 303-313
- PubMed: 8003966
- DOI: https://doi.org/10.1002/pro.5560030214
- Primary Citation of Related Structures:
1LTD - PubMed Abstract:
Flavocytochrome b2 from Saccharomyces cerevisiae catalyzes the oxidation of L-lactate to pyruvate and the electron transfer to cytochrome c in the mitochondrial intermembrane space. It is a homotetramer with a molecular weight of 4 x 58 kDa, each monomer of which is composed of 2 distinct domains, the one carrying FMN and the other, a "b5-like" heme. The native structure has been described at a resolution of 2.4 A (Xia ZX, Mathews FS, 1990, J Mol Biol 212:837-863). The heme domains protrude from the central body of the tetramer consisting of the 4 FMN binding domains. Because only 2 heme domains are visible in the electron density map, the other 2 are probably disordered. We crystallized the Escherichia coli recombinant flavocytochrome b2 from S. cerevisiae inhibited by sulfite. Although the crystals were obtained under very different conditions from those of the pyruvate-containing native enzyme, they were found to be isostructural (P 3(2) 2 1, a = b = 164.5 A, c = 114.0 A). The 2.6-A X-ray structure was extensively refined with X-PLOR (R = 17.3%), which made it possible to describe in detail the recombinant flavocytochrome b2 molecular structure. There exist few differences between the native and recombinant structures, in line with the fact that they show similar kinetic behavior, and they further confirm the intrinsic mobility of the heme domain (Labeyrie F, Beloil JC, Thomas MA, 1988, Biochim Biophys Acta 953:134-141). This structure will be used as a starting model in the structural resolution of flavocytochrome b2 point mutants.
Organizational Affiliation:
Istituto di Scienze Biochimiche, Università di Parma, Italy.