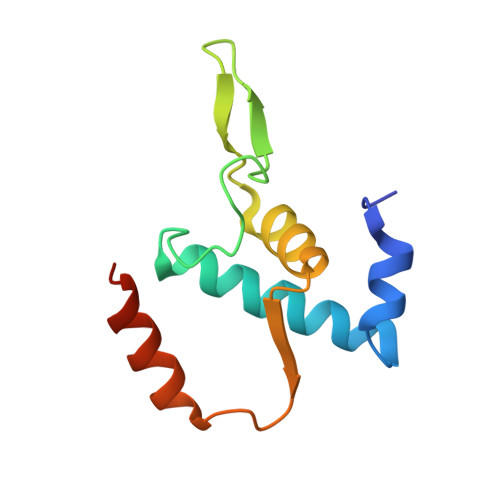
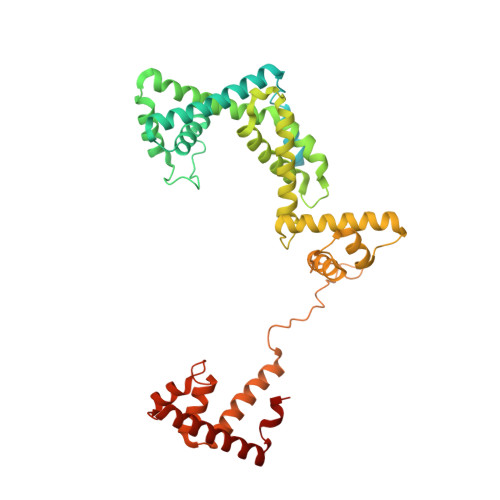
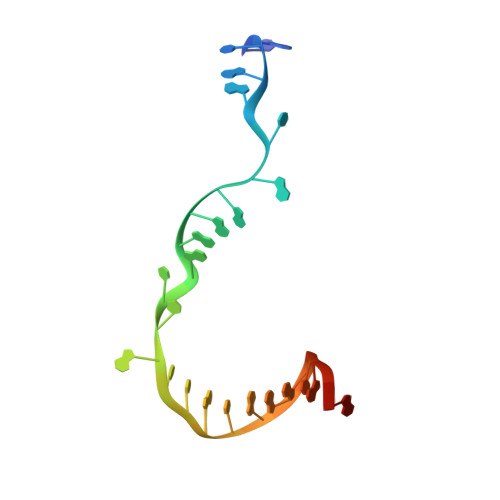
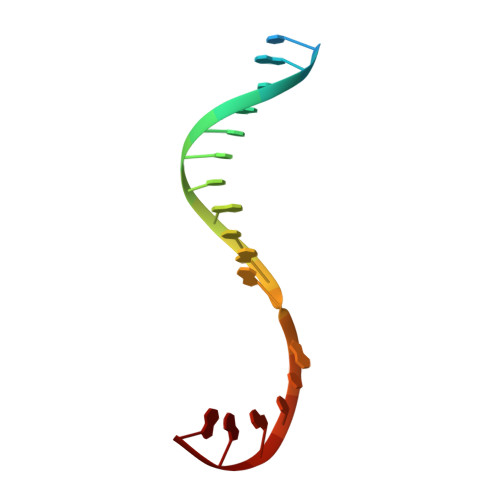
Promoter-sequence determinants and structural basis of primer-dependent transcription initiation in Escherichia coli .
Skalenko, K.S., Li, L., Zhang, Y., Vvedenskaya, I.O., Winkelman, J.T., Cope, A.L., Taylor, D.M., Shah, P., Ebright, R.H., Kinney, J.B., Zhang, Y., Nickels, B.E.(2021) Proc Natl Acad Sci U S A 118
- PubMed: 34187896
- DOI: https://doi.org/10.1073/pnas.2106388118
- Primary Citation of Related Structures:
7EH0, 7EH1, 7EH2 - PubMed Abstract:
Chemical modifications of RNA 5'-ends enable "epitranscriptomic" regulation, influencing multiple aspects of RNA fate. In transcription initiation, a large inventory of substrates compete with nucleoside triphosphates for use as initiating entities, providing an ab initio mechanism for altering the RNA 5'-end. In Escherichia coli cells, RNAs with a 5'-end hydroxyl are generated by use of dinucleotide RNAs as primers for transcription initiation, "primer-dependent initiation." Here, we use massively systematic transcript end readout (MASTER) to detect and quantify RNA 5'-ends generated by primer-dependent initiation for ~4 10 (~1,000,000) promoter sequences in E. coli The results show primer-dependent initiation in E. coli involves any of the 16 possible dinucleotide primers and depends on promoter sequences in, upstream, and downstream of the primer binding site. The results yield a consensus sequence for primer-dependent initiation, Y TSS-2 N TSS-1 N TSS W TSS+1 , where TSS is the transcription start site, N TSS-1 N TSS is the primer binding site, Y is pyrimidine, and W is A or T. Biochemical and structure-determination studies show that the base pair (nontemplate-strand base:template-strand base) immediately upstream of the primer binding site (Y:R TSS-2 , where R is purine) exerts its effect through the base on the DNA template strand (R TSS-2 ) through interchain base stacking with the RNA primer. Results from analysis of a large set of natural, chromosomally encoded E coli promoters support the conclusions from MASTER. Our findings provide a mechanistic and structural description of how TSS-region sequence hard-codes not only the TSS position but also the potential for epitranscriptomic regulation through primer-dependent transcription initiation.
Organizational Affiliation:
Department of Genetics, Rutgers University, Piscataway, NJ 08854.