Synthesis of RNA-cofactor conjugates and structural exploration of RNA recognition by an m6A RNA methyltransferase.
Meynier, V., Iannazzo, L., Catala, M., Oerum, S., Braud, E., Atdjian, C., Barraud, P., Fonvielle, M., Tisne, C., Etheve-Quelquejeu, M.(2022) Nucleic Acids Res 50: 5793-5806
- PubMed: 35580049
- DOI: https://doi.org/10.1093/nar/gkac354
- Primary Citation of Related Structures:
7P8Q, 7P9I, 7P9O - PubMed Abstract:
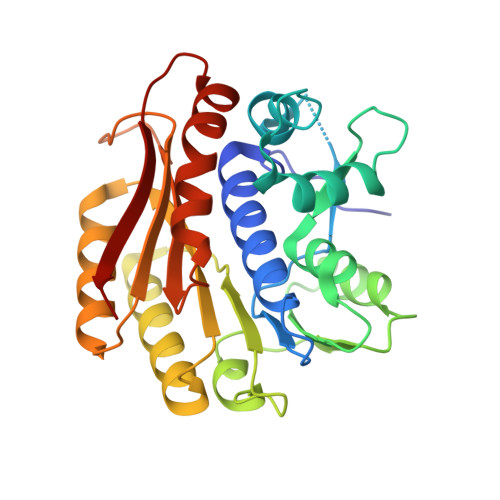
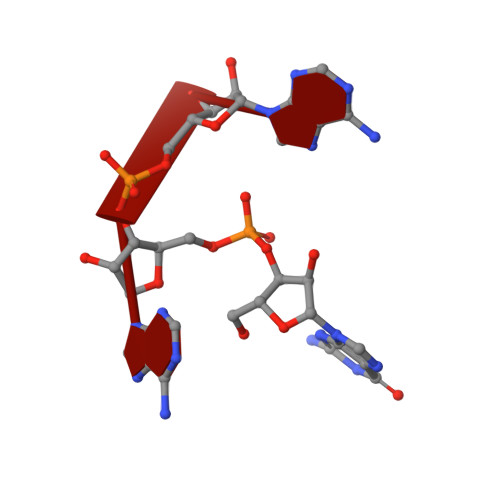
Chemical synthesis of RNA conjugates has opened new strategies to study enzymatic mechanisms in RNA biology. To gain insights into poorly understood RNA nucleotide methylation processes, we developed a new method to synthesize RNA-conjugates for the study of RNA recognition and methyl-transfer mechanisms of SAM-dependent m6A RNA methyltransferases. These RNA conjugates contain a SAM cofactor analogue connected at the N6-atom of an adenosine within dinucleotides, a trinucleotide or a 13mer RNA. Our chemical route is chemo- and regio-selective and allows flexible modification of the RNA length and sequence. These compounds were used in crystallization assays with RlmJ, a bacterial m6A rRNA methyltransferase. Two crystal structures of RlmJ in complex with RNA-SAM conjugates were solved and revealed the RNA-specific recognition elements used by RlmJ to clamp the RNA substrate in its active site. From these structures, a model of a trinucleotide bound in the RlmJ active site could be built and validated by methyltransferase assays on RlmJ mutants. The methyl transfer by RlmJ could also be deduced. This study therefore shows that RNA-cofactor conjugates are potent molecular tools to explore the active site of RNA modification enzymes.
Organizational Affiliation:
Expression Génétique Microbienne, UMR 8261, CNRS, Université Paris Cité, Institut de Biologie Physico-Chimique (IBPC),?75005,?Paris, France.